Professor, Plant & Microbial Biology
We are broadly interested in organelle formation in bacteria. We use the magnetosome organelles of magnetotactic bacteria as a model to understand the molecular mechanisms of compartmentalization and biomineral production. We are also beginning to explore the ferrosomes in some bacteria, and likely archaea, as a new model organelle.
Ph.D. Cell Biology UCSF, 2001
B.S. Biology MIT, 1996
Assembly and Subcellular Organization of Bacterial Organelles
Despite their apparent simplicity, bacterial cells are defined by a high degree of subcellular complexity and organization. One of the more dramatic examples of subcellular differentiation is the formation of protein- and lipid-bounded organelles by a variety of bacterial species. One such organelle, the magnetosome, is lipid-bounded and houses the machinery to build nano-sized magnetite crystals. We use magnetosomes as a model to understand the molecular mechanisms of compartmentalization and biomineral production. We are also beginning to explore the ferrosomes in some bacteria, and likely archaea, as a new model organelle.
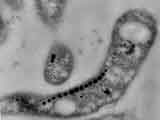
The magnetosome chains of magnetotactic bacteria are one of the best-studied examples of membranous bacterial organelles. Magnetosome chains (see figure above) contain 15-20 approximately 50-nm magnetite crystals that act like the needle of a compass to orient magnetotactic bacteria in geomagnetic fields, thereby simplifying their search for their preferred microaerophilic environments. The unique properties of magnetosomal magnetite crystals have drawn attention to their potential use in biotechnology, bioremediation, and geobiology and have made them a genetically tractable system for the study of biomineralization. In addition to these applications, the cell biological characteristics of magnetosomes make them ideal for the study of organelle biology in bacteria. Each magnetite crystal within a magnetosome is surrounded by a lipid bilayer, and specific soluble and transmembrane proteins are sorted to the magnetosome membrane. These results suggest that to build a magnetosome a bacterium must be able to generate a membranous comparmtent, target the appropriate set of proteins to this membrane and control their number and position within a cell.
My lab uses a combination of cell biological, genetic and biochemical approaches to define the physical characteristics of the magnetosome and identify key genes involved in controlling its production and function. Our earlier work showed that the magnetosome membrane is an independent organelle that pre-exists the formation of magnetite and that magnetite synthesis proceeds simultaneously from multiple adjacent magnetosomes. Recently, the work of several groups including ours has led to the identification of a large genomic region with many genes encoding proteins that are localized to the magnetosome and are essential for magnetite formation. The challenge now is to understand the specific functions of these genes and how their products interact to form a magnetosome.
More information at: http://www.komeililab.org
Follow us on twitter: @micromagnets and @MagnetoBerkeley
Rausser College of Natural Resources Distinguished Teaching Award - 2020
Fellow of the American Academy of Microbiology - 2019
Bakar Fellow - 2019
Miller Institute Faculty - 2015
Kavli Fellow - 2014
Winkler Family Fund - 2008
Hellman Family Fund - 2008
Fellowship in Science and Engineering - Packard Foundation - 2007
Junior Faculty Fellowship - Hellman Family Foundation - 2007

Arash Komeili
Berkeley, California 94720-3102